import duckdb
conn = duckdb.connect("embeddings.db")DuckDB, with version 1.0.0 released just last week, emphasizing API/format stability and backwards compatibility, is one of my favorite systems for doing data analysis. It’s blazingly fast, helps you avoid running out of memory even when dealing with larger datasets on a single machine and, quoting Hannes Mühleisen, one of its creators, it “doesn’t suck” when it comes to non-core database things like importing CSV or having a decent CLI. Last but not least, SQL is clearly undefeated:
Just in case you feel offended by this diagram: I’m a software engineer and I have trained and deployed generative language models with actual business value since 2021. So I’m in two circles here and I still think there’s a lot of truth in it.
Since I’m working on neural information retrieval use-cases, I was very excited when I heard that DuckDB has array support including distance functions like dot product and cosine and that it also got a vector similarity search extension recently. Equipped with these tools, we can use DuckDB to work with embeddings.
Embeddings
An embedding is a vector representation of data like a text or an image mapped into a vector space in some meaningful way, i.e. data points that are similar in some sense should be close to each other in that vector space. Embeddings are kind of a by-product of training neural networks, because they are an internal representation of the data inside the model. But we can also use their properties for search or recommendation: if we’re able to obtain useful embeddings, we can search for similar content, or for documents that are similar to a query, by searching for documents with similar embedding vectors.
I don’t want to go into more details here about embeddings. If you want to learn more, there are plenty of good explanations. Two articles that I found helpful are
- What are embeddings? by Vicky Boykis.
- Embeddings: What they are and why they matter by Simon Willison.
Instead, I want to show you how to store embeddings along with their documents in DuckDB, how to integrate with an embedding model running locally, and how to do vector search with DuckDB.
The examples are in Python, so if you want to follow along, install the required dependencies:
pip install duckdb transformers FlagEmbedding polarsWe start by creating a database on disk and connecting to it:
Loading Data into DuckDB
In order to do vector search, we need some content first, so let’s download some documents and store them in our database. To keep things simple, we’ll use the pre-processed Wikipedia dataset provided by the Wikimedia Foundation as a set of Parquet files. To test the multilingual capabilities of our embedding model, we’ll use the German Wikipedia subset 20231101.de. You can get a nice preview of the data using the Hugging Face dataset viewer:
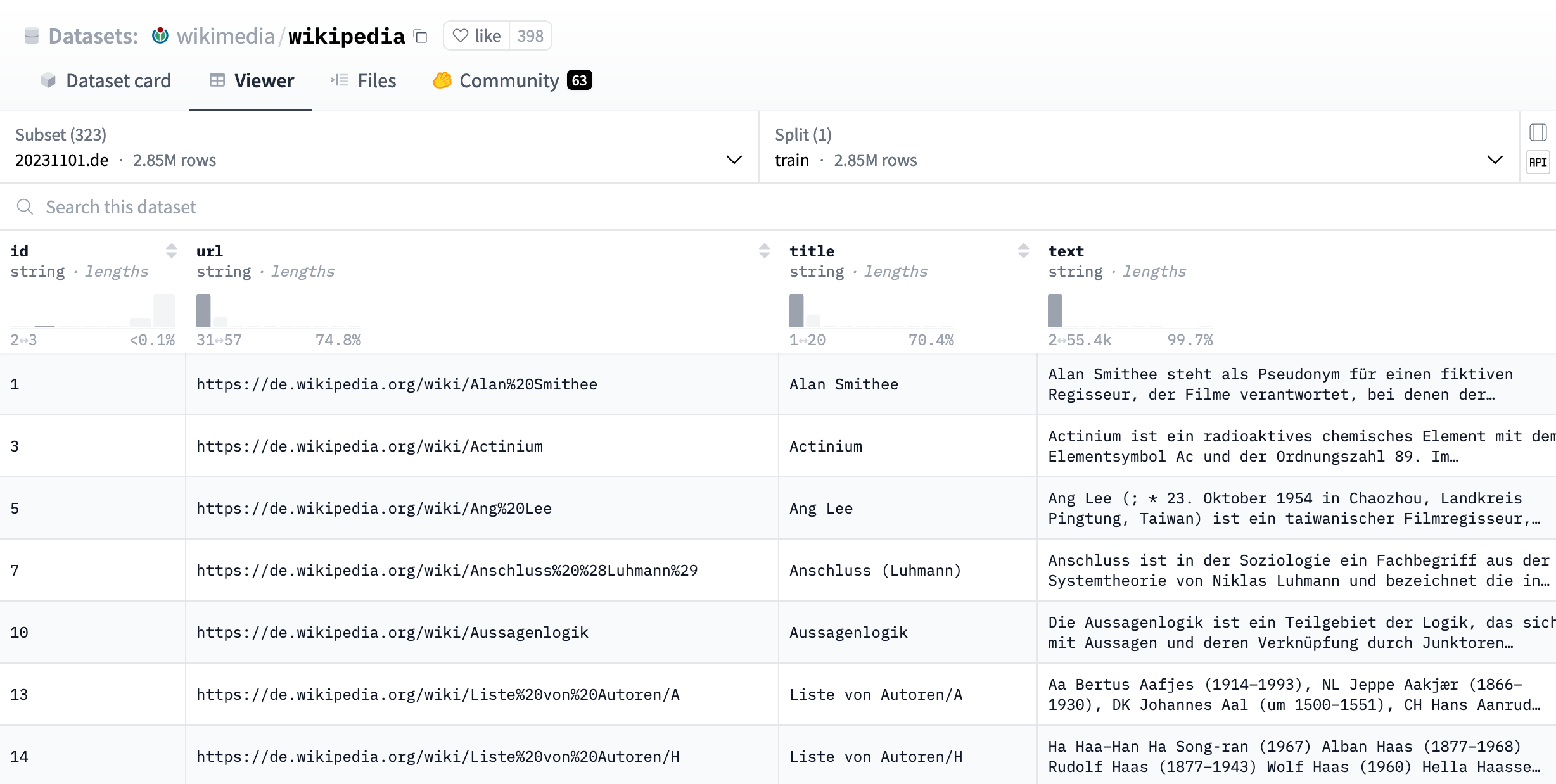
We can use the new DuckDB integration with Huggingface Datasets that was just announced to fetch the data directly from the hub. Let’s try to verify the 2.85M rows the viewer reports for the German subset with a DuckDB query:
FROM 'hf://datasets/wikimedia/wikipedia/20231101.de/*.parquet'
SELECT count(*) AS count;| count |
|---|
| 2845308 |
The count looks close enough. I was surprised though that it took only a few seconds to run this query on my machine, even though the German Wikipedia subset is more than 5GB compressed data spread over 20 Parquet files. How is that possible?
The reason is that DuckDB supports partial reading of Parquet files, which means that if your query states that you only need a part of the data (filter or projection), or even just metadata, DuckDB can optimize the reads and only read what it really needs from disk. Even more amazingly, it also works with Parquet files served over HTTP if your webserver supports HTTP range requests. Read more about how DuckDB can efficiently perform queries on Parquet in this blog post.
Let’s fetch the first 10 entries to have a quick look:
FROM 'hf://datasets/wikimedia/wikipedia/20231101.de/*.parquet'
SELECT * LIMIT 10;| id | url | title | text |
|---|---|---|---|
| "1" | "https://de.wikipedia.org/wiki/… | "Alan Smithee" | "Alan Smithee steht als Pseudon… |
| "3" | "https://de.wikipedia.org/wiki/… | "Actinium" | "Actinium ist ein radioaktives … |
| "5" | "https://de.wikipedia.org/wiki/… | "Ang Lee" | "Ang Lee (; * 23. Oktober 1954 … |
| "7" | "https://de.wikipedia.org/wiki/… | "Anschluss (Luhmann)" | "Anschluss ist in der Soziologi… |
| "10" | "https://de.wikipedia.org/wiki/… | "Aussagenlogik" | "Die Aussagenlogik ist ein Teil… |
| "13" | "https://de.wikipedia.org/wiki/… | "Liste von Autoren/A" | " Aa Bertus Aafjes (1914–199… |
| "14" | "https://de.wikipedia.org/wiki/… | "Liste von Autoren/H" | " Ha Haa–Han Ha Song-ran (19… |
| "15" | "https://de.wikipedia.org/wiki/… | "Liste von Autoren/C" | " Ca Cab–Cap Fernán Caballe… |
| "16" | "https://de.wikipedia.org/wiki/… | "Liste von Autoren/I" | " I Yi I (1536–1584) Ia I… |
| "17" | "https://de.wikipedia.org/wiki/… | "Liste von Autoren/K" | " Ka Dieter B. Kabus (1941–19… |
Note that this also only takes a few seconds for the reason explained above. Depending on your query, it could of course take longer as DuckDB has to fetch more data.
Finally, let’s create a table directly from this query, with a few more articles:
CREATE TABLE wikipedia AS
FROM 'hf://datasets/wikimedia/wikipedia/20231101.de/*.parquet'
SELECT * limit 200;Now we have 200 entries in our local database which means we can query them much faster:
SELECT text FROM wikipedia LIMIT 5;| text |
|---|
| "Alan Smithee steht als Pseudonym für einen fiktiven Regisseur, der Filme verantwortet, bei denen der… |
| "Actinium ist ein radioaktives chemisches Element mit dem Elementsymbol Ac und der Ordnungszahl 89. I… |
| "Ang Lee (; * 23. Oktober 1954 in Chaozhou, Landkreis Pingtung, Taiwan) ist ein taiwanischer Filmregi… |
| "Anschluss ist in der Soziologie ein Fachbegriff aus der Systemtheorie von Niklas Luhmann und bezeich… |
| "Die Aussagenlogik ist ein Teilgebiet der Logik, das sich mit Aussagen und deren Verknüpfung durch Ju… |
Loading an Embedding Model
In order to create our vectors for our documents, we need an embedding model. While we could use an embedding as a service provider like Cohere, OpenAI or Google, I want to use an open model that we can run locally without any subscription.
BGE-M3 from Chen et al. is an interesting choice due to its multi-lingual capabilities and that it can produce not only dense embeddings, but also multi-vector term embeddings for late interaction as popularized by ColBERT. We can even ask the model to output sparse vectors to help with traditional TF/IDF or BM-25 like search with an inverted index. In this example we’re just using dense, single vector embeddings though.
The BGE-M3 weights are available on Hugging Face as well and we can load the model as follows:
from FlagEmbedding import BGEM3FlagModel
import torch
device = "cpu"
# use a GPU if available to speed up the embedding computation
if torch.cuda.is_available(): device = "cuda" # Nvidia GPU
elif torch.backends.mps.is_available(): device = "mps" # Apple silicon GPU
model = BGEM3FlagModel('BAAI/bge-m3', use_fp16=True, device=device)We can run the model on a few inputs, get the dense embeddings and compute similarities with the dot product:
queries = ["What is BGE M3?", "What is DuckDB?"]
documents = [
"BGE M3 is an embedding model supporting dense retrieval, lexical matching and multi-vector interaction.",
"DuckDB is a fast in-process analytical database. It supports a feature-rich SQL dialect complemented with deep integrations into client APIs",
]
query_embeddings = model.encode(queries)["dense_vecs"]
document_embeddings = model.encode(documents)["dense_vecs"]
similarity = query_embeddings @ document_embeddings.T
similarityarray([[0.626 , 0.1918],
[0.3362, 0.732 ]], dtype=float16)Creating Embeddings for Wikipedia Articles
Now that we have a few documents and a model, we can create embeddings for each document and store them in our database as well. We create an embeddings table with two columns: One to store our vectors we get from our model, float arrays of size 1024 in this case. And another for the document id to link back to our table with the Wikipedia pages:
CREATE TABLE embeddings(
doc_id VARCHAR,
embedding FLOAT[1024]
);There are multiple ways to create our embeddings. For example, we could
- Loop through batches of inputs in Python, call the embedding model and store each batch in the database.
- Create a custom DuckDB function (UDF) to call the model and write the embeddings in a single SQL statement.
- Use another library like Hugging Face Datasets for batch processing and then write the results to our database.
Looping through Arrow Batches
Let’s go with the first option here: Fetch batches as Arrow records via the DuckDB Apache Arrow integration and write back batches of embeddings. We’ll see an example of the second option later when we create a UDF to compute the query embedding. First, we need to create a reader that will return an Arrow RecordBatch iterator from our query result, with each batch containing 100 records:
reader = conn.execute(
"FROM wikipedia SELECT id, text WHERE id NOT IN (FROM embeddings select doc_id);"
).fetch_record_batch(100)We can iterate over these batches and call the model to compute embeddings for each batch. Note that here we are reading batches of size 100 from the database into main memory, and then smaller batches of size 4 into GPU memory (if our model runs on the GPU). Once we have a batch of 100 embeddings, we put them into an Arrow table, write them back to the database using the Arrow import feature, and fetch the next batch. We also convert our embeddings back to float32 because DuckDB doesn’t currently support 16 bit floats.
import pyarrow as pa
import numpy as np
for batch in reader: # 100 records per batch
# 4 records per GPU batch
embeddings = model.encode(batch["text"].tolist(), batch_size=4)["dense_vecs"].astype(np.float32)
# add our embeddings to the batch
batch = batch.add_column(0, "embedding", list(embeddings))
# we need an Arrow table for the DuckDB replacement scan to work
batch_table = pa.Table.from_batches([batch])
conn.cursor().execute(
"INSERT INTO embeddings FROM (FROM batch_table SELECT id, embedding);")You will probably want to run this on a GPU for more than a few dozen documents, or it will take a long time. You may also want to change batch_size depending on the size of your GPU memory.
Let’s have a look at our embeddings:
FROM wikipedia JOIN embeddings ON (wikipedia.id = embeddings.doc_id)
SELECT *
LIMIT 5;| text | embedding |
|---|---|
| "Actinium ist ein radioaktives chemisches Element mit dem Elementsymbol Ac und der Ordnungszahl 89. I… | [0.026413, -0.018646, -0.0625, 0.037933, -0.010872, … 0.013496] |
| "Ang Lee (; * 23. Oktober 1954 in Chaozhou, Landkreis Pingtung, Taiwan) ist ein taiwanischer Filmregi… | [-0.067383, -0.025024, -0.012299, 0.01944, -0.008987, … -0.01368] |
| "Anschluss ist in der Soziologie ein Fachbegriff aus der Systemtheorie von Niklas Luhmann und bezeich… | [-0.030182, 0.01799, -0.018402, 0.031464, -0.013596, … 0.020737] |
| "Die Aussagenlogik ist ein Teilgebiet der Logik, das sich mit Aussagen und deren Verknüpfung durch Ju… | [0.007038, 0.014084, -0.012436, 0.007843, -0.010979, … -0.058868] |
| " Ha Haa–Han Ha Song-ran (1967) Alban Haas (1877–1968) Rudolf Haas (1877–1943) Wolf Haas (1960) He… | [-0.066101, 0.001437, -0.023911, -0.034119, 0.011597, … -0.013344] |
DuckDB being a real database gives us a lot of flexibility to store our embeddings and join them back with the documents or chunks they are generated from. This is even more powerful if we take into account the numerous data source extensions. While we’re reading from remote Parquet files here, we could just as easily fetch our data from Postgres using DuckDB’s Postgres reader, a CSV in some bucket, or even a combination of different sources.
Running Vector Similarity Search
We’re almost there! Now we just need to embed our queries as well before we can do a similarity search. To do this, let’s see another DuckDB feature in action: We’ll create a user-defined function (UDF) embed using the Python Function API to call our model:
from duckdb.typing import VARCHAR
def embed(sentence: str) -> np.ndarray:
return model.encode(sentence)['dense_vecs']
conn.create_function("embed", embed, [VARCHAR], 'FLOAT[1024]')Once we’ve registered the function with DuckDB, we can use in like any native function as part of our SQL query:
SELECT embed('Who was the first human on the moon?') AS query_embedding;| query_embedding |
|---|
| [0.011353, 0.010292, -0.058899, -0.044098, 0.015778, … -0.033691] |
Equipped with our embedding UDF, we can use a array distance function like array_cosine_similarity, or just array_inner_product for normalized vectors to find the most similar documents to our query based on their embedding vectors. Let’s define a small search function that given a query, returns the most similar Wikipedia articles:
def search(q: str):
return conn.execute("""
FROM embeddings JOIN wikipedia ON (wikipedia.id = embeddings.doc_id)
SELECT wikipedia.id, title, array_inner_product(embedding, embed($q)) AS similarity
ORDER BY similarity DESC
LIMIT 5""",
{"q": q}
).pl()
search('Who was the first human on the moon?')| id | title | similarity |
|---|---|---|
| "207" | "Apollo-Programm" | 0.591756 |
| "256" | "Apollo 11" | 0.576521 |
| "22" | "Liste von Autoren/B" | 0.485747 |
| "26" | "Liste von Autoren/M" | 0.481897 |
| "16" | "Liste von Autoren/I" | 0.465602 |
Here we can see the capabilities of a multilingual model like BGE-M3 in action. Even though our query is in English and our documents are German, articles with relevant information to answer the question have a high similarity. If we ask the question in German as well, the result is almost the same.
search('Wer war der erste Mensch auf dem Mond?')| id | title | similarity |
|---|---|---|
| "207" | "Apollo-Programm" | 0.608182 |
| "256" | "Apollo 11" | 0.602915 |
| "26" | "Liste von Autoren/M" | 0.483409 |
| "16" | "Liste von Autoren/I" | 0.458147 |
| "22" | "Liste von Autoren/B" | 0.450969 |
Note that the model was trained on Wikipedia (among others) so this would not be a good dataset for actually validating how well the model performs. The interesting question you have to ask yourself is how well it works, in your domain, on your data.
Using an HNSW Index
By default DuckDB has to compare a query embedding with all document embeddings to find the most similar ones. With our small sample of 200 documents that’s not an issue, but if you’re trying to find the most similar ones among millions of documents, comparing the query vector with each document vector will become expensive in terms of computation and latency.
To speed up vector search (at the cost of some accuracy), we can use an approximate nearest neighbor (ANN) method. A popular approach for ANN is the Hierarchical Navigable Small World (HNSW) algorithm. DuckDB supports HNSW through its vector similarity search extension.
Let’s load the vss extension and create an index on our embedding table:
INSTALL vss;
LOAD vss;
-- required for persistent databases as of DuckDB 1.0.0
SET GLOBAL hnsw_enable_experimental_persistence = true;
CREATE INDEX ip_idx ON embeddings USING HNSW (embedding)
WITH (metric = 'ip');Now we can use the index for our vector search. Note that currently, the index is only used when our similarity search looks like in the top_k common table expression, so we can’t combine it i.e. with a WHERE clause for filtering or a join immediately. We can use the results to filter and join as we like though:
q = "Who was the first human on the moon?"WITH top_k AS (
FROM embeddings SELECT *
ORDER BY array_inner_product(embedding, embed($q))
LIMIT 5)
FROM top_k JOIN wikipedia ON (wikipedia.id = top_k.doc_id)
SELECT wikipedia.id, title, array_inner_product(embedding, embed($q)) AS similarity
ORDER BY similarity DESC| id | title | similarity |
|---|---|---|
| "207" | "Apollo-Programm" | 0.591756 |
| "256" | "Apollo 11" | 0.576521 |
| "22" | "Liste von Autoren/B" | 0.485747 |
| "26" | "Liste von Autoren/M" | 0.481897 |
| "16" | "Liste von Autoren/I" | 0.465602 |
The result looks still good, but, depending on your HNSW index settings, accuracy will not be as good as comparing all vectors. This trade-off might be necessary to achieve faster performance though.
We can also check if the index is used by putting EXPLAIN statement in front of the SQL query:
EXPLAIN
SELECT *
FROM embeddings
ORDER BY array_inner_product(embedding, embed('some text'))
LIMIT 5;┌───────────────────────────┐
│ PROJECTION │
│ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ │
│ #0 │
│ #1 │
└─────────────┬─────────────┘
┌─────────────┴─────────────┐
│ PROJECTION │
│ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ │
│ embedding │
│ doc_id │
│ NULL │
└─────────────┬─────────────┘
┌─────────────┴─────────────┐
│ HNSW_INDEX_SCAN │
│ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ │
│embeddings (HNSW INDEX SCAN│
│ : ip_idx) │
│ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ │
│ embedding │
│ doc_id │
│ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ ─ │
│ EC: 5 │
└───────────────────────────┘
Since HNSW is experimental for persistent databases as of DuckDB 1.0.0, let’s remove the index again or we might run into WAL issues when loading the database from disk next time.
DROP INDEX ip_idx;Conclusion
This article has demonstrated the power and flexibility of DuckDB for handling embeddings and performing vector search. We’ve leveraged DuckDB’s efficient Parquet handling to load data from large datasets, integrated a PyTorch embedding model seamlessly using both batch processing and UDFs, and performed vector similarity search with and without an HNSW index.
The ability to store embeddings along with your data directly in DuckDB, coupled with its extensive data source connectivity, opens up a wide range of possibilities for complex analysis and retrieval.
We have barely scratched the surface of what is possible with DuckDB, and there is much more to explore. In an upcoming article, we are going to work with slightly more complex vector operations and see if we can do simple ColBERT-style late interaction by implementing the MaxSim operator using only native DuckDB functions.
I hope you enjoyed reading this article. If you have have questions, corrections or any other feedback, don’t hesitate to contact me through any of the channels linked on the about page.